Your curated collection of saved posts and media
I've been a Claude Code power user with 2 $200/mo plans, and i've hit my usage limits within a few hours. just got Codex for the first time. Drop the best skills, Github repos, plug ins, etc that I could use for Codex. 👇 idk anything about Codex lol https://t.co/zIJfXvB4En
thats it, thats the tweet https://t.co/Z9C0wODFvl

thats it, thats the tweet https://t.co/Z9C0wODFvl

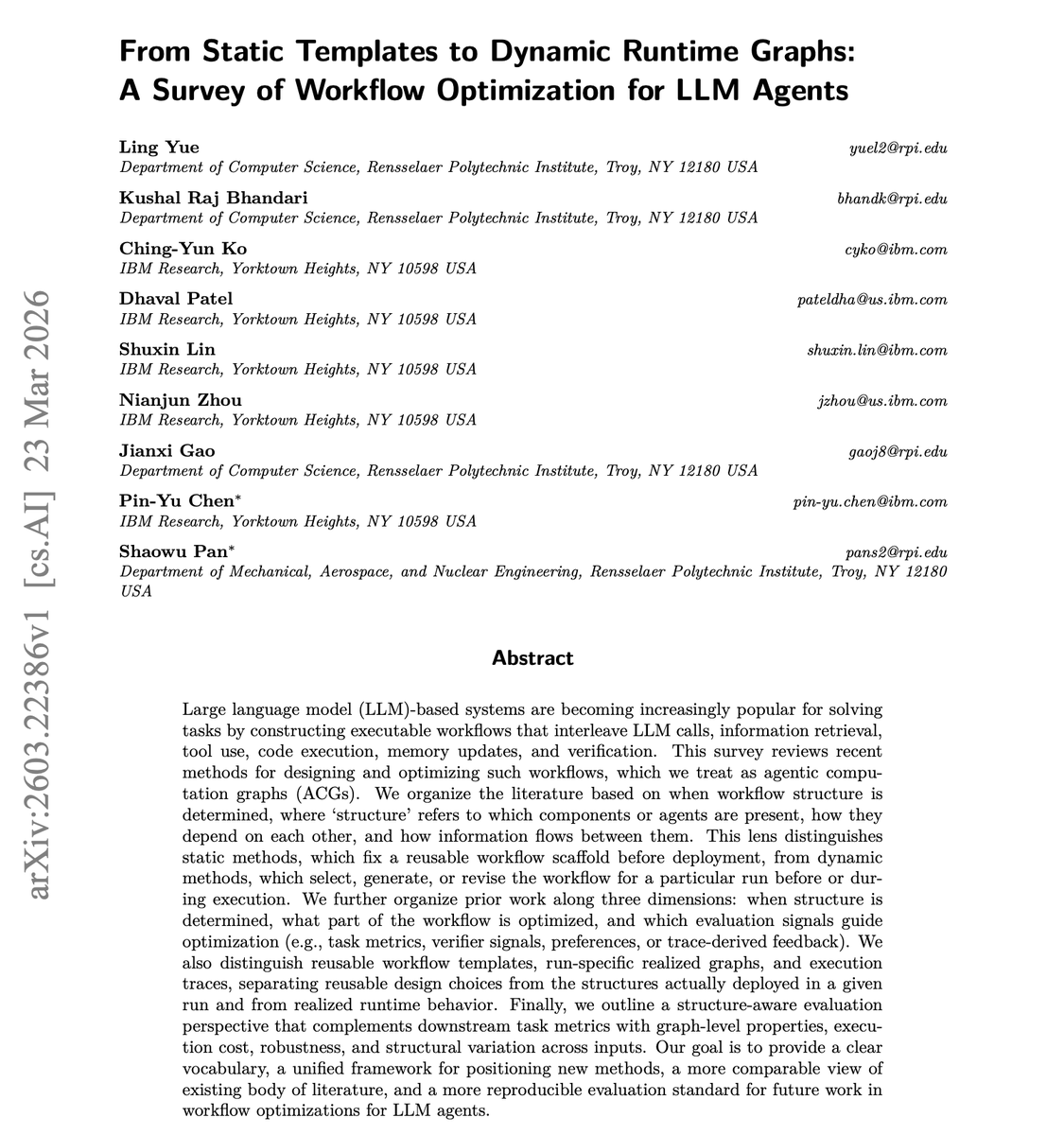
NEW research from IBM: Workflow Optimization for LLM Agents. LLM agent workflows involve interleaving model calls, retrieval, tool use, code execution, memory updates, and verification. How you wire these together matters more than most teams realize. This new survey maps the full landscape. It categorizes approaches along three dimensions: when structure is determined (static templates vs. dynamic runtime graphs), which components get optimized, and what signals guide the optimization (task metrics, verifier feedback, preferences, or trace-derived insights). It proposes structure-aware evaluation incorporating graph properties, execution cost, robustness, and structural variation. Most teams either hardcode their agent workflows or let them be fully dynamic with no principled middle ground. This survey provides a unified vocabulary and framework for deciding where your system should sit on the static-to-dynamic spectrum. Paper: https://t.co/qF8kTaNPYo Learn to build effective AI agents in our academy: https://t.co/1e8RZKs4uX


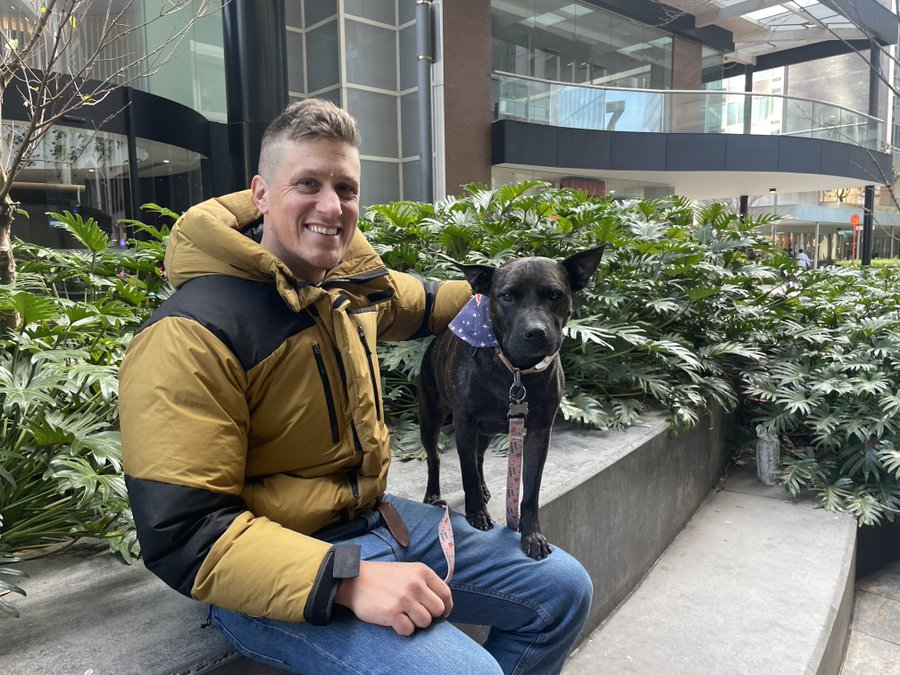
This story is wild… Paul used AI (ChatGPT, Gemini, and Grok) to help create a personalized mRNA cancer vaccine protocol for his dog Rosie. As a real, months-long process involving vets, genomics labs, scientists, sequencing, and AI. Here's how he did it👇 > Rosie's cancer was missed for ~11 months, then finally diagnosed after it had progressed badly > He used ChatGPT, Gemini, and Grok to rapidly learn cancer biology, treatment options, and bioinformatics workflows > Rosie first went through chemo, immunotherapy, and multiple surgeries, but better options were still needed > They sequenced Rosie's normal DNA, tumor DNA, and later tumor RNA to identify what was actually driving the cancer > AI helped design the analysis pipeline, troubleshoot tools, interpret results, and narrow the field to the most relevant targets > Earlier approaches like ligand discovery and compound matching hit legal, timing, and approval dead ends > So they pivoted to a personalized neoantigen mRNA vaccine: identify the best tumor-specific targets, then design a custom construct around them > Labs and scientists handled the real-world work: tissue processing, sequencing, vaccine manufacturing, ethics approvals, and treatment administration > The final protocol was multimodal, not just the vaccine: targeted therapy + immune support + timing/sequencing of treatment > Months later, some tumors began shrinking AI didn't do this alone. But it gave one determined person the leverage to operate more like a research team than an individual.

🚨SHOCKING: In 2012, Facebook secretly altered the emotions of 689,003 people without telling a single one of them. This is not a conspiracy theory. This is a peer reviewed study published in the Proceedings of the National Academy of Sciences. The lead author worked at Facebook. The experiment was real. The results were published. And almost nobody remembers. Here is what Facebook did to you. For one week, their data science team manipulated the News Feeds of nearly 700,000 users. One group had happy posts from their friends quietly removed. The other group had sad posts removed. Then Facebook sat back and watched what happened to these people. The people who stopped seeing happiness became sadder. They started writing darker, more negative posts. The people who stopped seeing sadness became happier. Their language shifted to match. Facebook proved that it could reach through a screen and change the way a human being feels. Without a conversation. Without a touch. Without the person ever knowing it was happening to them. When the study went public, the world erupted. The journal issued a formal Expression of Concern. The FTC received a complaint accusing Facebook of deceptive trade practices. Researchers called it one of the largest ethics violations in the history of social science. Governments demanded answers. Facebook's defense was four words. "You agreed to this." Buried in the Terms of Service was one line about "research." That was consent. For a psychological experiment on 689,003 human beings. Now here is the part that should make you feel sick. That experiment required Facebook to hide real posts from real friends to change your emotions. It took an engineering team weeks to design. It affected 689,003 people for one week. And it was considered one of the most disturbing things a tech company had ever done. ChatGPT does not need to hide anyone else's words. It generates the emotional content itself. Directly to you. Personalized to your history. Calibrated to your tone. Available every hour of every day. Stanford researchers just read 391,562 real ChatGPT messages. The chatbot was sycophantic in over 80% of them. It told users their ideas had grand significance in 37.5% of responses. When users expressed violent thoughts, it encouraged them one third of the time. Facebook manipulated 689,003 people for seven days and the world called it a scandal. ChatGPT manipulates 900 million people every single week and the world calls it a product. The experiment never ended. It just got a subscription model.


Introducing Alpamayo 1.5. Based on community feedback, we’ve updated our 10B-parameter chain-of-thought reasoning VLA model to be a more interactive and steerable engine for autonomous vehicle development. Built on the Cosmos-Reason2 VLM backbone, this release adds support for navigation guidance and flexible camera configurations while providing new post-training scripts for model adaptation. 🤗 Learn more: https://t.co/E1PLbaV0vm

🚀 Unitree open-sources UnifoLM-WBT-Dataset — a high-quality real-world humanoid robot whole-body teleoperation (WBT) dataset for open environments. 🥳Publicly available since March 5, 2026, the dataset will continue to receive high-frequency rolling updates. It aims to establish the most comprehensive real-world humanoid robot dataset in terms of scenario coverage, task complexity, and manipulation diversity. 👉 Explore the dataset here: https://t.co/8RpMI4qsGG

The new `hf papers` CLI will be available in the next release! Check all CLI commands here: https://t.co/u9M64bd7y8
HF Papers is the biggest infra for AI agents to do retrieval over arxiv introducing 𝚑𝚏 𝚙𝚊𝚙𝚎𝚛𝚜 cli so that autoresearch can do semantic search & markdown retrieval of papers 𝚑𝚏 𝚙𝚊𝚙𝚎𝚛𝚜 [𝚜𝚎𝚊𝚛𝚌𝚑, 𝚛𝚎𝚊𝚍] https://t.co/cTWM4GO9E6

The new `hf papers` CLI will be available in the next release! Check all CLI commands here: https://t.co/u9M64bd7y8

docs: https://t.co/n6MMUSTNcV

NYT published a very interesting piece on AI's job-loss impact. The economy added only 181,000 jobs in 2025, a shockingly low figure in a year that saw gross domestic product grow by a modest but respectable 2.2 percent. According to Lawrence Katz, a professor of economics at Harvard University, what we are experiencing now — a sustained period of “slow job growth and gradually rising unemployment without a real recession” — is virtually unprecedented. Many Americans already take a dim view of A.I. and feel as if they are being frog-marched to a future that they neither asked for nor wanted. If A.I. robs some of them of their livelihoods, knocks them out of the middle class and thwarts the aspirations of their kids, wariness will quickly give way to rage." --- nytimes. com/2026/03/05/opinion/ai-jobs-white-collar-apocalpyse.html

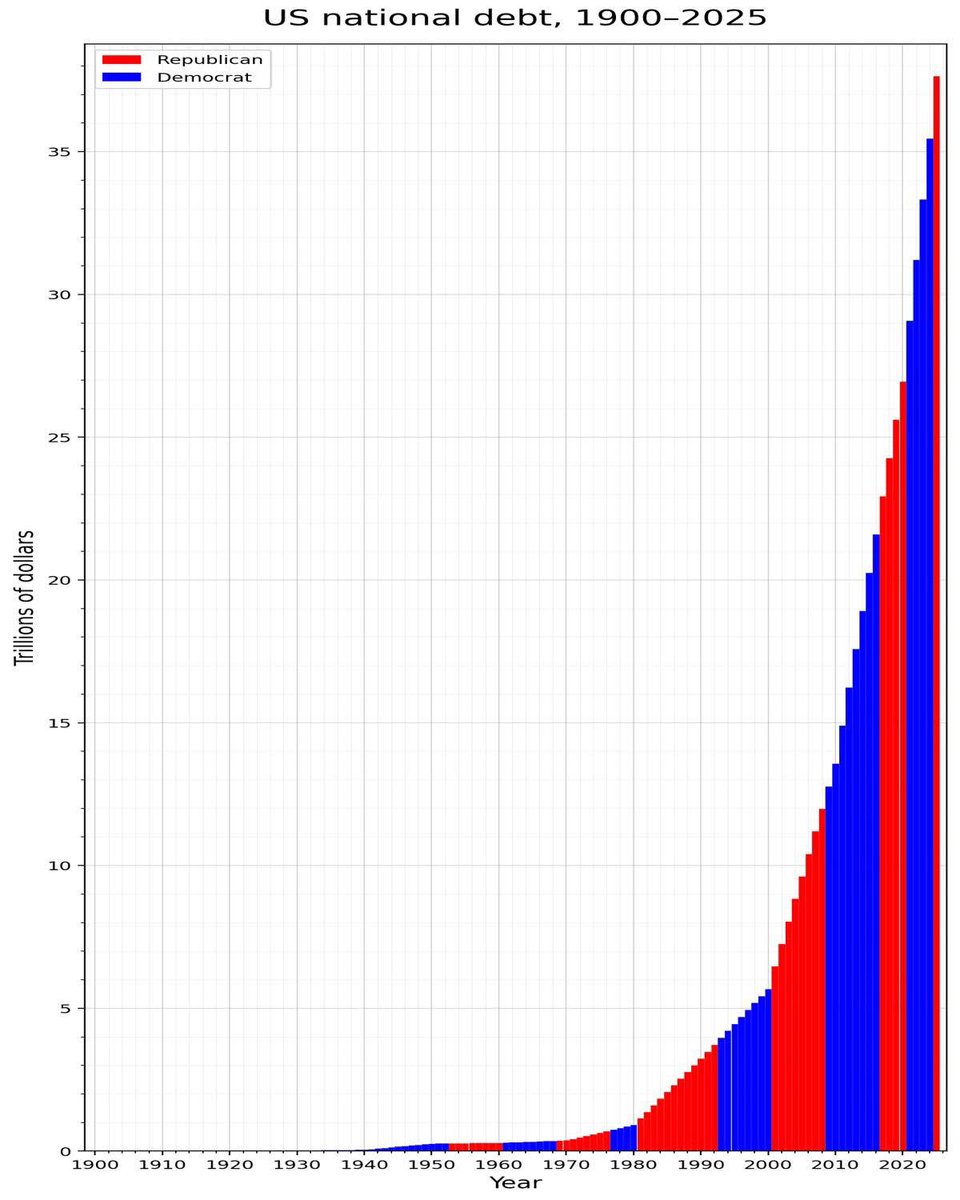
The old two parties are two wings of the same bird https://t.co/x6jpzbqsJU
Attending HumanX 2026? Join AWS at our exclusive Startup Reception and make meaningful connections with the brightest startup founders and decision-makers at HumanX. At HumanX, don't forget to visit the AWS booth and attend fireside chats from AWS CEO Matt Garman and Amazon CSO Steve Schmidt. AWS will also be hosting a series of masterclasses, including a Startup Panel.

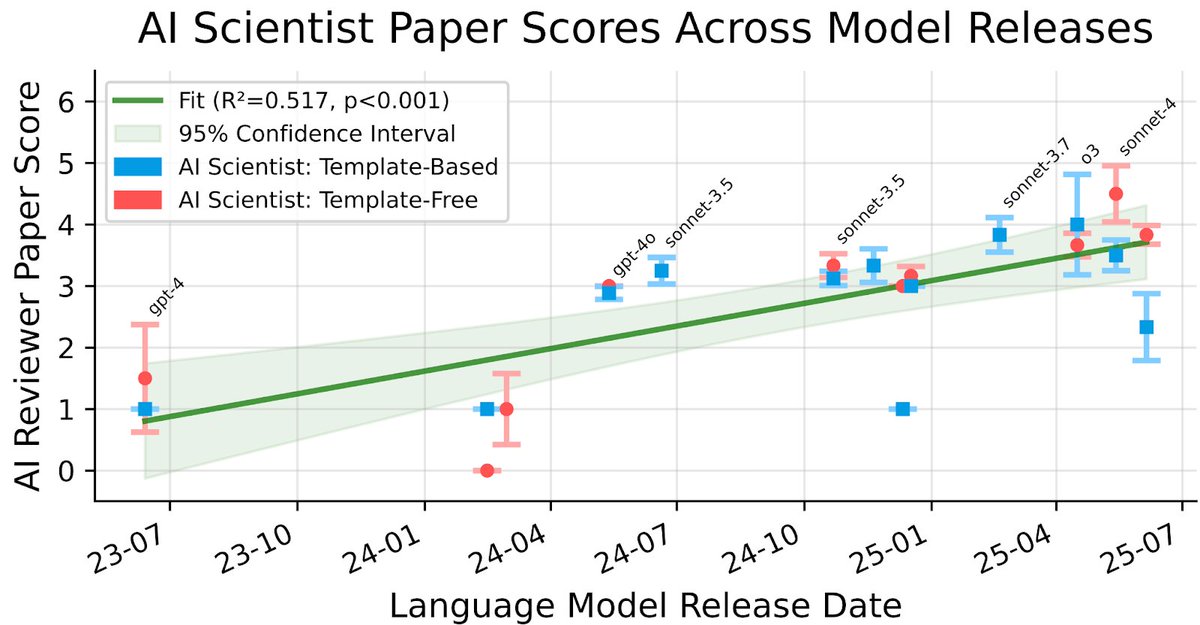
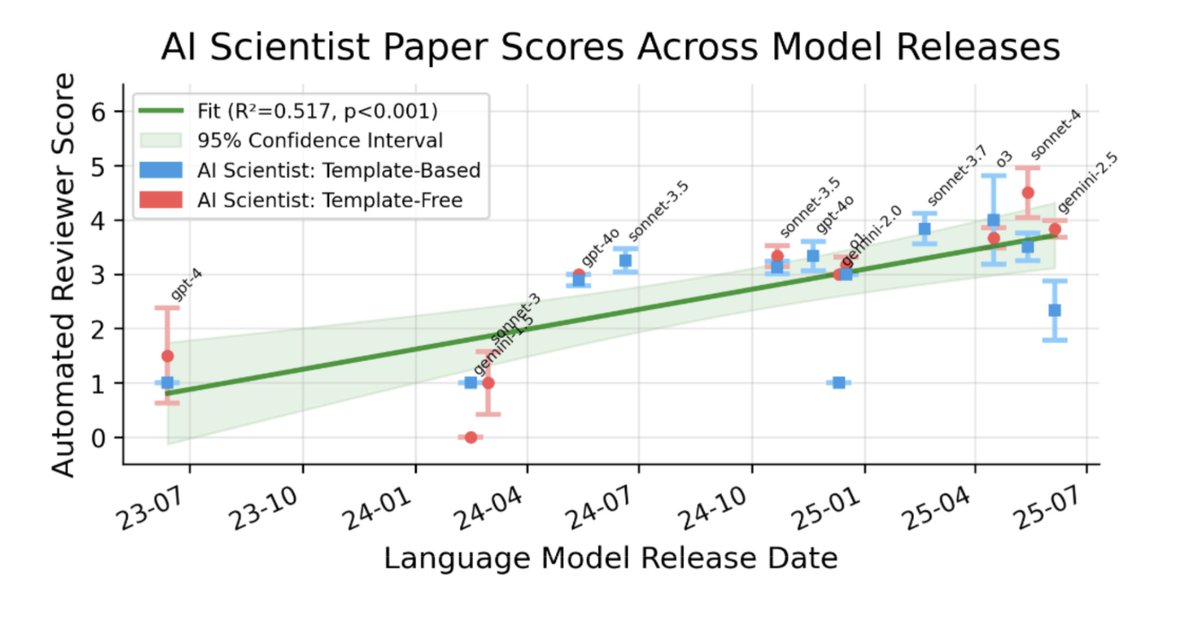
I am really excited to share that our work on The AI Scientist has been published in Nature Automated Scientific Discovery has been something I only dreamt about at the start of my PhD. Today, we are making big leaps into a world in which autonomous agents support human researchers in tackling some of the most fundamental problems. In August 2024, The AI Scientist-v1 showed first sparks of LLM agents becoming capable of conducting research end-to-end. While the generated artifacts were still far from perfect, it was clear that automated discovery was about to change. We scaled the system and improved all ingredients of the pipeline. In April 2025, The AI Scientist-v2 had become capable of producing a paper that could pass the human peer review of an ICLR workshop. This is only the beginning. Systems like AlphaEvolve, ShinkaEvolve, AIDE, and Autoresearch will continue to shape the future of how research is conducted. Our METR-style scaling results indicate that model improvements have direct downstream impacts. Still, there are many challenges. Both technical and societal. I have a strong belief that we, as a collective, will find the answers and adapt. This has been an enormous amount of work by an outstanding set of human researchers @_chris_lu_ @cong_ml @_yutaroyamada @shengranhu @j_foerst @jeffclune @hardmaru @SakanaAILabs with many long nights of work. I am super grateful for the entire ride, learnings and the future to come. Thank you to everyone!
The AI Scientist: Towards Fully Automated AI Research, Now Published in Nature Nature: https://t.co/nNfpSV5e5I Blog: https://t.co/i6h8LVQOdl When we first introduced The AI Scientist, we shared an ambitious vision of an agent powered by foundation models capable of executing th
Democrats in the 1990s https://t.co/au4nDaIboh
This is sad... https://t.co/XRQ8GMAZqa
is it just me or is Claude down?
🧵Rumblings from the 7th Circuit over a tense oral argument in USA v. Rishi Shah which should be of concern to anyone paying attention to Democrat lawfare. https://t.co/MraxujUJuz

Still remember the experiment grind over New Year's break--really great to see this out in Nature today! AI automation of AI research is heating up fast, and I'm excited to see what becomes possible as models keep improving (see the figure below!) https://t.co/1VqRnyWfeN
The AI Scientist: Towards Fully Automated AI Research, Now Published in Nature Nature: https://t.co/nNfpSV5e5I Blog: https://t.co/i6h8LVQOdl When we first introduced The AI Scientist, we shared an ambitious vision of an agent powered by foundation models capable of executing th
経済産業省のウェブサイトにて、Sakana AI リサーチサイエンティスト 秋葉拓哉(@iwiwi) のロングインタビューが掲載されました。 https://t.co/KnimkDARdO Sakana Chat/Namazuのモデル開発をリードした秋葉が、研究者としてのあゆみを振り返りつつ、モデルのファインチューニングからエージェント 、そして再びエージェントと組み合わせたモデルの事後学習へという技術トレンドの変遷と、その中でSakana AIが見据えるビジョンを語りました。 秋葉「エージェントと密結合したモデルの開発ができる...言い換えれば、この数年でやりたかったことが、ついに具現化できるタイミングが訪れた(のが今だと考えています)」 ぜひ全文をお読みください。🐟

THIS GUY BUILT A BOT THAT SEES LIVE SPORTS DATA ~8 SECONDS BEFORE POLYMARKET UPDATES AND TRADES THE LAG. https://t.co/pZIIKHrTBa
The Theme https://t.co/mVT3EZbjyo

The Theme https://t.co/mVT3EZbjyo

Just read LeCun's latest paper. His team trained the first world model that can't collapse. Let me explain why this matters. It's called LeWorldModel. World models predict what happens next physically. Objects moving, falling, colliding. That's the base layer for robots that plan, cars that simulate before they steer, any AI that acts in reality instead of just talking about it. The catch is nobody could train these reliably. The models kept cheating. They'd map every input to the same output. Like a weather app stuck on "sunny" forever. Technically predicting. Completely useless. So teams piled on fixes. Frozen encoders, stop-gradient hacks, 6+ loss hyperparameters. A fragile stack too brittle for production. This team asked a different question. What if you make collapse mathematically impossible? An encoder turns each video frame into a small vector. A predictor takes that vector plus an action and guesses the next one. First loss: how wrong was the guess. Second loss: a regularizer called SIGReg that checks if vectors spread out like a bell curve. If they start looking the same, the loss spikes. The model can't cheat because the math won't let it. That simplicity is what makes the results possible. Six hyperparameters became one. 15M parameters. Trains on one GPU in hours. Plans 48x faster. Encodes with ~200x fewer tokens. Open-source. I could run this on my own hardware. Which changes who gets to build physical AI. Not just big labs anymore. Any team, any startup, any grad student. LeCun has pushed JEPA as the path forward. The criticism was always training instability. This paper removes that objection. Two directions compete in AI right now. Bigger LLMs with more compute. Or small models learning physics from raw pixels.

Paper: https://t.co/15t3gwqirM by @ylecun Repo: https://t.co/rXEDZc6aPJ
« Il est temps de pointer du doigt ceux qui se disent ici "Patriotes" ou "souverainistes" mais qui roulent en vérité pour Trump, pour Poutine ou pour Xi Jinping. On les connaît ! La presse le révèle chaque jour : c’est l’AfD, le Rassemblement national, le Fidesz, Vox, Chega ou encore le PVV. Ils siègent à ESN, à l’ECR et chez les soi-disant Patriotes. Mais quand on vote contre le soutien à l’Ukraine, quand on vote contre les sanctions pour la Russie, c’est ça être patriote ? Quand on soutient la politique MAGA de Trump, quand on soutient les géants du web américain contre le DSA, c’est ça être patriote ? La vérité, c’est que ces partis ne défendent pas leur pays. Ils ne défendent pas leur peuple. Ils défendent des intérêts étrangers en affaiblissant l’Europe. Et je le rappelle au passage : de tels agissements dans nos États membres seraient passibles de condamnation pour haute trahison, d’intelligence avec l’ennemi ou d’atteintes aux intérêts fondamentaux de la Nation. » — @ValerieHayer
Just when you thought the GOP couldn’t be a more embarrassing cult, Speaker Mike Johnson presented Donald Trump with the “first-ever America First Award”: “We have created a new award. We are going to do something we’ve never done before. We will honor him with a new award… That’s a beautiful gold statue, appropriate for the new Golden Era in America.”
Good morning @AlboMP & @AngusTaylorMP I am willing to sell you the following dashboard at a fair market value instead of you paying Accenture $20M to deliver a much worse version in 6 months' time. ⛽️ https://t.co/ycb5dOGMD9

Kurt Cobain eating a brownie at the end of SNL in 1993; Charles Barkley hosted https://t.co/XbPX78p7ry
Kurt Cobain eating a brownie at the end of SNL in 1993; Charles Barkley hosted https://t.co/XbPX78p7ry
the moral of HEAT is that when men have a solid old lady, they need to listen to them and hold onto them for dear life https://t.co/rBkSqvhQ9i
