@BoWang87
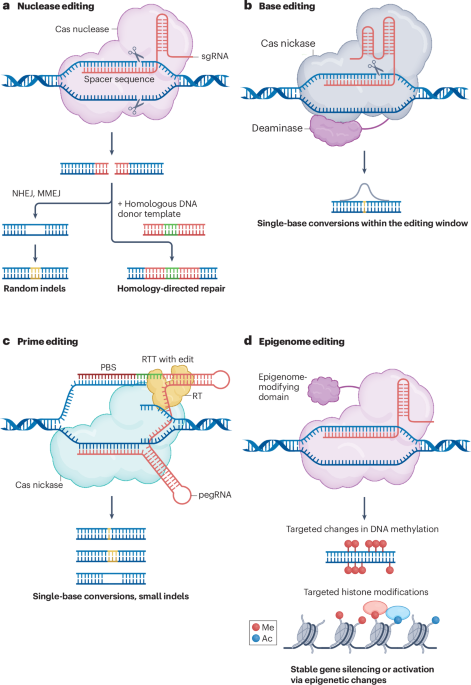
Thrilled to share our review paper, out today in @NatureRevGenet : "Harnessing artificial intelligence to advance CRISPR-based genome editing technologies" Full paper : 🔗 https://t.co/ZBJcgDZduY CRISPR has already changed medicine. AI is now changing CRISPR. We spent a long time mapping the full landscape of where machine learning and deep learning are having real, measurable impact across the genome editing workflow — and where the most exciting opportunities lie ahead. Here's what we cover: Guide RNA design — Deep learning models now predict on- and off-target activity for Cas9, Cas12, Cas13, and emerging systems like TnpB and IscB. We've gone from sequence heuristics to transformer-based models that generalize across organisms. Cell-type-specific generalization remains a frontier. Base and prime editing — ML models predict bystander effects, product purity, and editing efficiency from sequence context alone. For prime editing, tools like PRIDICT and DeepPE have made pegRNA design far more tractable at scale. Enzyme engineering — Protein language models (ESM, EVOLVEpro) are now guiding directed evolution of Cas proteins — expanding PAM compatibility, reducing immunogenicity, improving compactness — at a pace impossible through classical lab iteration alone. Novel enzyme discovery — Foundation models trained on metagenomics are uncovering entirely new CRISPR systems from microbial diversity: new Cas variants, TnpB systems, and eukaryotic Fanzor proteins. The search space is enormous; AI is how we navigate it. Virtual cell models — This is where I'm most excited. AI-powered virtual cells can, in principle, predict the functional consequences of any edit in any cell type — selecting targets, anticipating off-targets, modeling tissue-specific outcomes. But realizing this vision requires causally-rich, contextually diverse perturbation data. Scale of data matters as much as scale of model. Delivery — ML-guided LNP design is closing the last mile between an edit that works in a dish and one that works in a patient. Across all of this, one theme recurs: AI accelerates where data is abundant and well-structured. The field's next challenge is generating that data at the right diversity and scale. This paper was a true collaboration. Huge thanks to Tyler Thomson, Gen Li, Amy Strilchuk, @HAOTIANCUI1 , and Bowen Li — you each brought something irreplaceable to this. Special shoutout to @BowenLi_Lab for his leaderhsip in this work!