@s_batzoglou
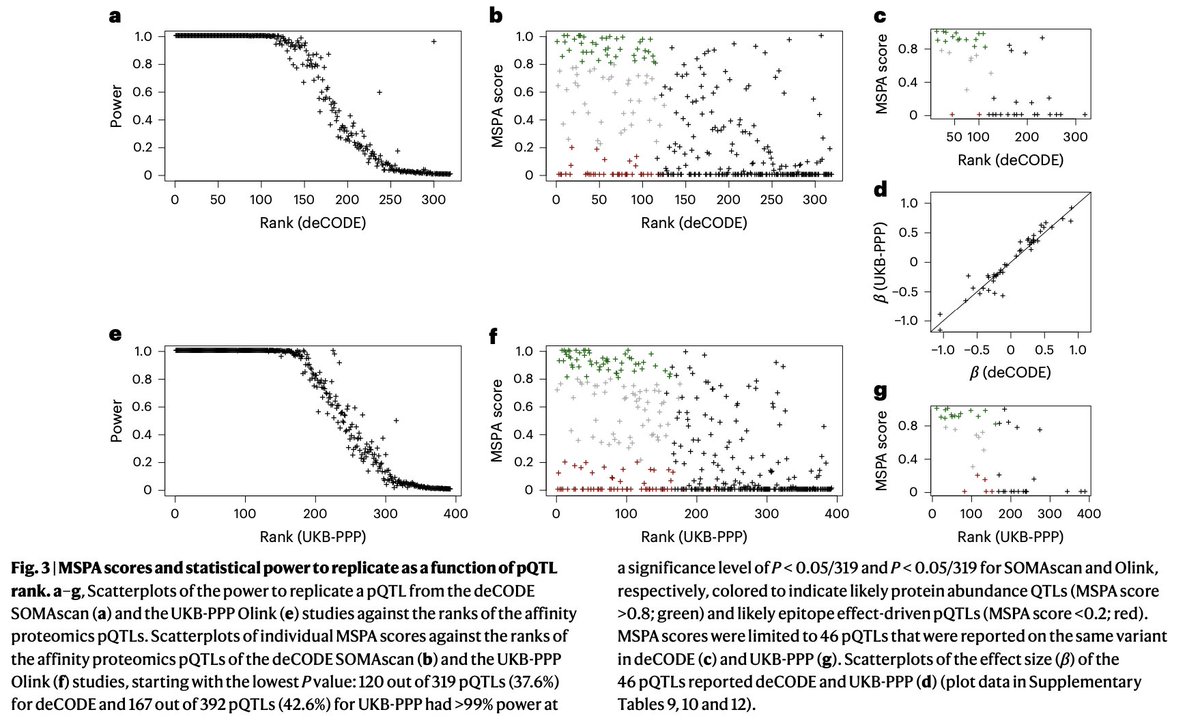
A new proteogenomic study by @ksuhre , performing pQTL GWAS on a discovery and a validation cohort using @seer_bio nanoparticle enrichment and mass spectrometry. Also, pQTLs found in the UK Biobank by Olink and in the deCODE Icelandic cohort by SomaScan are evaluated for their evidence in the Seer MS data (when present). About 30% show strong evidence, and 30% show weak to no evidence. Among the evaluated pQTLs found by both Olink and SomaScan, which are likely more reliable, almost all show strong evidence. This implies that the scoring method using MS data is informative on the reliability of affinity-based pQTLs.